|
12/29/2023 0 Comments Jmol gfp protein databank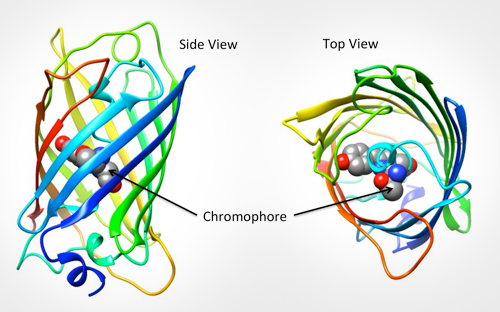
A clear distinction between sequences capable of forming a chromophore and those that do not have the residues required for chromophore formation can be obtained by examining a single measurement-the RMSD of the overlap of the central alpha helices of the crystal structure of S65T GFP and the AlphaFold2 determined structure. Statistical analysis of these measurements showed that both programs conclusively distinguished between chromophore forming and non-chromophore forming proteins. The resultant structures were mined for a series of geometric measurements that are crucial to chromophore formation. The structures of 21 sequences of GFP-like fluorescent proteins that will post-translationally form a chromophore and of 23 GFP-like non-fluorescent proteins that do not have the residues required to form a chromophore were determined by AlphaFold2 and RoseTTAfold. Their training has not only taught them protein structure and folding, but also chemistry. Our genetic analysis supports conclusions from structural and biochemical studies and demonstrates a critical role for multiple, highly conserved glycine residues in GFP protein stability.ĪlphaFold2 and RoseTTAfold are able to predict, based solely on their sequence whether GFP-like proteins will post-translationally form a chromophore (the part of the protein responsible for fluorescence) or not. Many mutations in this category generate substitutions of highly conserved glycine residues, including Gly67 in the chromogenic tripeptide Gl圓1, Gl圓3, and Gl圓5 in the second β-strand and Gly20, Gly91 and Gly127 in the lids of the β-barrel scaffold. By contrast, other loss-of-fluorescence mutations substantially diminish the amount of GFP protein, suggesting that they compromise protein stability. Mutations that alter residues important for chromophore maturation, such as Arg96 and Ser205, greatly reduce or extinguish fluorescence without dramatically altering GFP protein accumulation. Here we report a collection of ethyl methansulfonate-induced GFP loss-of-function mutations in the model plant Arabidopsis thaliana. However, loss-of-function mutations that would aid in understanding GFP protein folding and chromophore formation have not been fully catalogued. Random and site-specific mutagenesis has been used to optimize GFP fluorescence and create derivatives with novel properties. The GFP chromophore is generated spontaneously in the presence of oxygen by a multi-step reaction involving cyclization of the internal tripeptide Ser65 (or Thr65)-Tyr66-Gly67, which is embedded in the center of an eleven-stranded β-barrel structure. Green fluorescent protein (GFP) and related fluorescent proteins are widely used in biological research to monitor gene expression and protein localization in living cells. The cavities of highlighter proteins that can undergo cis/trans isomerization have chemospatial properties that are common to both cis and trans GFP-like proteins. (5) The chemospatial characteristics of the chromophore cavity determine the isomerization state of the chromophore. These chromophoric deviations from planarity play an important role in determining the fluorescence quantum yield. (4) In most GFP-like proteins the protein matrix exerts a significant strain on planar chromophores forcing most GFP-like proteins to adopt non-planar chromophores. (3) The structural prerequisites for light activated proton pumps exist in many structures and it's possible that like bioluminescence, proton pumps are secondary functions of GFP-like proteins. (2) The primary/ancestral function of GFP-like proteins may well be to aid in light induced electron transfer. They are important to the function of GFP-like proteins, perhaps in protecting the chromophore or in β-barrel formation. The conclusions reached in this review are that (1) The lid residues are highly conserved, particularly those on the "top" of the β-barrel. We have used the pdb to examine the function of fluorescent proteins (FP) in nature, aspects of excited state proton transfer (ESPT) in FPs, deformation from planarity of the chromophore and chromophore maturation. This is the first systematic analysis of all the GFP-like structures in the pdb. The RCSB protein databank contains 266 crystal structures of green fluorescent proteins (GFP) and GFP-like proteins.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
